On February 6, BGI-Research, in collaboration with Fudan University, made a significant advance in metabolic disease research. The accomplishment was published online in Cell Metabolism today. By utilizing BGI Group’s spatial-temporal multi-omics technology, Stereo-seq, and its high-throughput single-cell nuclear sequencing technology, this pioneering study mapped nearly half a million cells in the hypothalamus of the macaque (Macaca fascicularis), creating the world's first primate hypothalamus cellular atlas of diabetes. This study offers new insights into the intricate workings of the hypothalamus and its relationship with prevalent disorders such as diabetes and obesity.
 The study, Region-specific transcriptomic responses to obesity and diabetes in macaque hypothalamus, is published in Cell Metabolism.
The study, Region-specific transcriptomic responses to obesity and diabetes in macaque hypothalamus, is published in Cell Metabolism.
Diabetes and obesity represent two of the most challenging metabolic disorders in the world, with their prevalence increasing at an alarming rate. The hypothalamus, a higher neural center regulating visceral and endocrine functions, is now at the forefront of research for its role in glucose and lipid metabolism. As the control center for the body's homeostasis, it has become a key subject for diabetes and obesity research.
Dr. Lei Ying, the first author of this paper and a researcher at BGI-Research, stated, "This study provides a comprehensive single-cell and spatial atlas of gene expression for the macaque hypothalamus, revealing the transcriptional level changes due to obesity and diabetes. It not only helps us to better understand the mechanisms behind metabolic diseases but also provides clues for discovering new drug targets. We hope this research will lead to more accurate and effective treatment strategies for obesity and diabetes."
In the study, researchers observed that diabetes is associated with stronger transcriptional changes across various cell types in the hypothalamus than obesity. Obesity primarily affects the infundibular nucleus (INF), which is involved in feeding, metabolism, fertility, and cardiovascular regulation. Diabetes, however, extends its impact to the paraventricular nucleus (PVN), which is associated with a myriad of functions including osmoregulation, appetite, and stress response through different hormones.
Distinct molecular changes in response to diabetes were observed between the INF and PVN. In a diabetes status, the INF showed increased inflammatory responses, whereas the PVN exhibited suppressed cellular metabolism and activity. These findings provide direct molecular evidence pinpointing the INF-PVN circuit as crucial in maintaining energy balance and highlight the differential responses of these two nuclei in developing targeted treatments for metabolic disorders.
The study also identified significant genes and pathways in the macaque hypothalamus that may be involved in metabolic regulation in primates. For instance, the study found that there was no expression change of HTR2C in POMC neurons in obese macaques, but there was an increase in AGRP neurons compared to normal macaques, suggesting that HTR2C signaling in AGRP neurons could be a critical target for controlling obesity.
HTR2C is a type of serotonin receptor highly expressed in POMC neurons. Activating this receptor can excite POMC neurons, thereby suppressing food intake, according to previous studies.
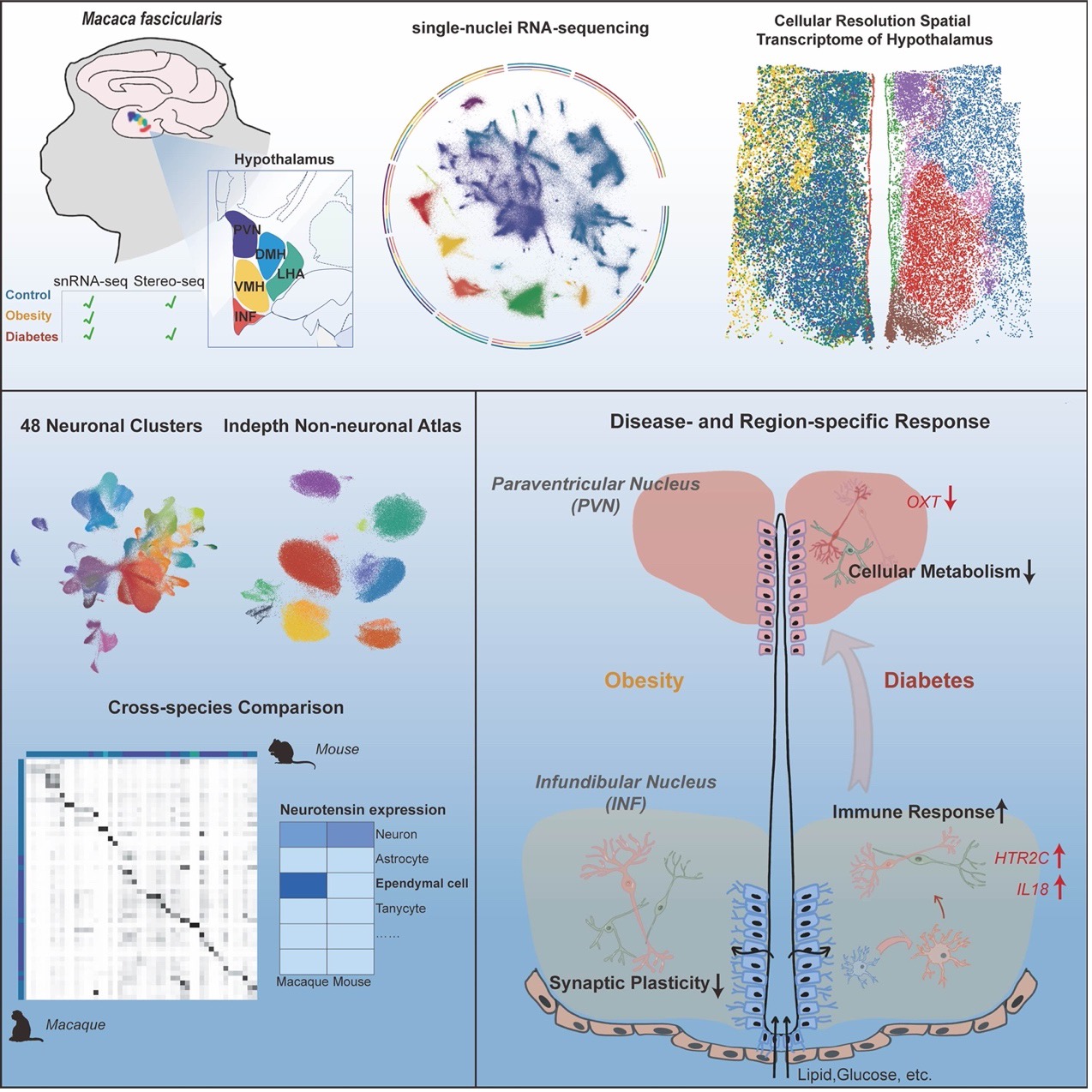 Key findings of the study.
Key findings of the study.
Another critical finding is the expression of Neurotensin (NTS), a multifunctional neuropeptide related to energy metabolism, which was initially identified within the neurons of rodents. In this study, researchers found for the first time that in macaques, NTS expression is not confined to neurons but is also significantly present in ependymal cells; in mice, however, it was expressed only in neurons. Interestingly, this primate-unique NTS population of ependymal cells is also present in humans. The reduction of neurotensin during obesity status may imply that NTS acts within the brain as an intercellular regulatory factor, playing a potential role in metabolic regulation.
The article can be read here: https://doi.org/10.1016/j.cmet.2024.01.003



