The iconic platypus and echidna (monotremes) display a unique mix of mammalian and reptilian features and form the most distantly related group of living mammals. One of their most well-known features is that they lay eggs to produce offspring while secret milk to feed them. Monotremes also have a complex sex chromosome system consisting of five pairs of sex chromosomes, while other mammals have only one pair of sex chromosome. As this clade is the sister group of therian mammals, it occupies an important position in mammalian evolution thus serves as a key to our understanding of the origin of mammals.
 Article link: https://www.nature.com/articles/s41586-020-03039-0
Article link: https://www.nature.com/articles/s41586-020-03039-0
Today, a paper presented the genomes of these egg-laying mammals, platypus and echidna, was published in the Nature journal. This large project involved over 40 researchers from institutes such as BGI-Shenzhen, University of Copenhagen, Zhejiang University, the University of Adelaide, and produced and analysed the first ever echidna genome and a greatly improved, chromosome-level platypus genome sequence. Prof. Guojie Zhang from BGI-Shenzhen, University of Copenhagen and Kunming Institute of Zoology, Prof. Qi Zhou from Zhejiang University and Prof. Frank Grutzner from the University of Adelaide share the corresponding authors of the paper.
Mammalian evolution
Human shares the most recent common ancestor with platypus and echidna about 180 million years ago. Comparing the monotreme and human genomes can inform us about the ancestral genomic structure of all mammals and genomic changes during early mammalian evolution. Thanks to the high-quality monotreme assemblies, for the first time, the team successfully reconstructed the mammalian ancestral karyotype as 2n=60 chromosomes. This provides important information to the study of genomic changes during early mammal evolution.
The platypus and echidna are the only egg-laying mammals, thus provide the key to understanding the change in the transition of reproductive strategy from egg-laying to production of live young in all other mammals. The team found that both monotremes keep one of the three vitellogenin genes, which encode the major egg proteins that is responsible for making the yolk in chicken, while these genes have become lost in other mammals. Genes involved in the milk production process such as casein genes were found to be present in both monotremes, suggesting that they have been evolved in the common ancestor of all mammals.
Adult monotremes have no teeth and their stomach are poorly developed. Correspondingly, the team found that both species have lost many of the key genes that are important in the development of teeth and stomach. These include odontogenic ameloblast-associated gene which participates in tooth development process such as tooth initiation and generation, and gastrin gene which stimulates gastric acid secretion and induce insertion the ATPase pumping H+ into stomach.
The system to safely remove oxygen carrier, the haemoglobin from blood, was previously thought to be common among all mammals. However, the key gene haptoglobin involved in the detoxification process is found missing in both platypus and echidna genomes. This revealed that the systems are different between monotreme and therians, and the system of human is most likely to arise quite recently in the ancestor of marsupial and eutherian mammals.
Although platypus and echidna both belong to the monotreme lineage and have diverged for only about 55 million years, they live in distinct environments. Platypuses are semi-aquatic, prey through electrolocation and feed on small invertebrates in fresh water. Echidnas live on land and hunt for termites via their well-developed olfactory system. The number of olfactory receptors is found to be considerably larger in echidna, while the number of vomeronasal receptors is larger in platypus. This is consistent with the different size of their olfactory bulbs and accessary olfactory bulb. There are fewer bitter taste receptors in monotremes compared to other mammals. These gene patterns may result from the adaptation to different environments in these species.
Sex chromosome evolution
“More than 15 years ago we discovered that monotremes have different sex chromosomes to all other mammals” says Prof. Frank Grutzner from the University of Adelaide, who co-led the study. “This finding revolutionised our understanding of mammalian sex chromosome evolution but also raised fundamental new questions. With the new genomes we can now tackle important questions about how sex chromosomes control monotreme sexual development and reproduction, which will be relevant for monotreme conservation and captive breeding programs”, he explains.
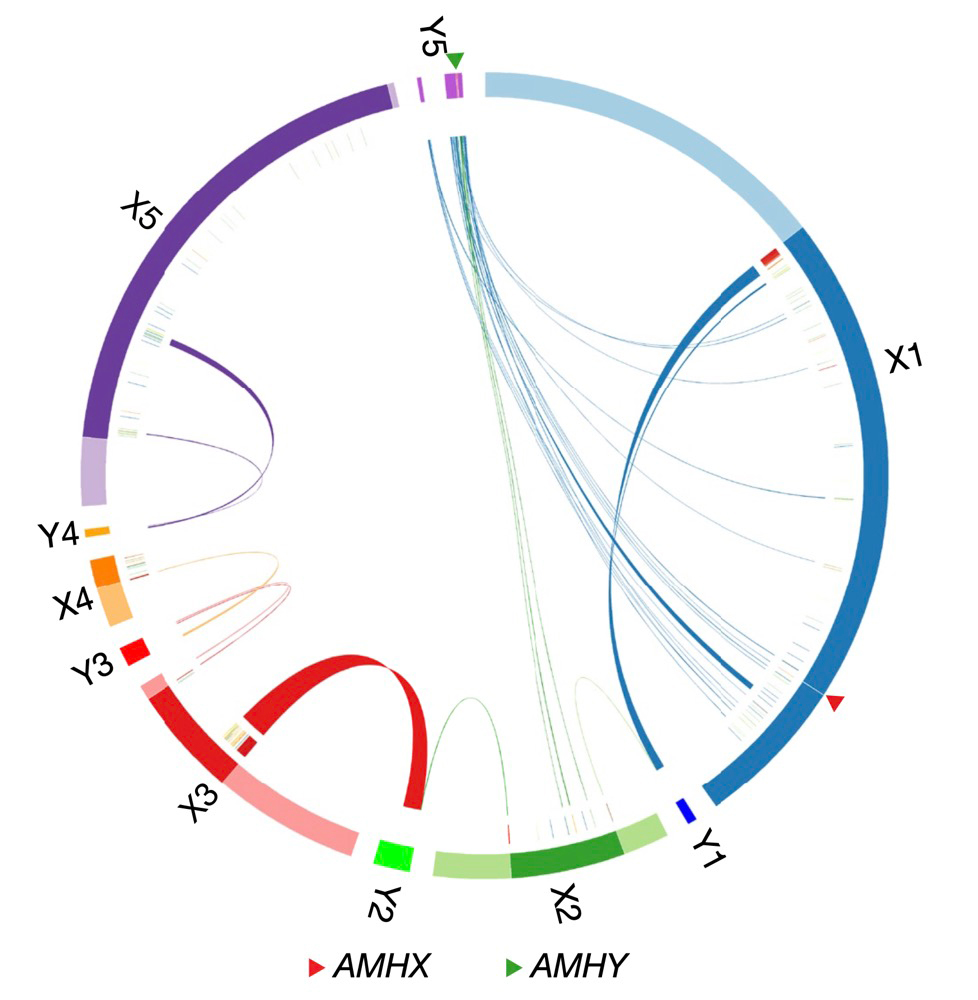
Monotremes’ X chromosomes share no homology with human’s X but partly with chicken’s Z. By the comparison among different species, the team found that monotreme’s five pairs of sex chromosomes are more likely to result from multiple reciprocal translocation events. “We found that in platypus X1 is the oldest sex chromosome, followed by X2, X3 and X5 and X4 is the youngest.” says Prof. Qi Zhou from Zhejiang University. “We were surprised that most genes on X1 are homologous with genes on Y5 instead of Y1, which is paired with X1 during meiosis. This implies that the platypus sex chromosomes had probably formed a ring structure in its early evolution. We also found unexpected inter-chromosomal interactions between the sex chromosomes in monotremes. This may account for the evolution of multiple sex chromosomes system in monotremes.”
“With the new sequencing technologies, we are able to obtain higher quality chromosome-level genome assemblies. Sex chromosome is long-term challenge for genome assembly, but we were able to solve the issue with the new method that we established” says Prof. Guojie Zhang from BGI-Shenzhen and University of Copenhagen, one of the corresponding authors of the paper. “And high-quality genomes of the key species in evolution can help us understand the genomic evolution process and the underlying genomic mechanisms of evolutionary innovations and adaptation” he adds.



