In a remarkable demonstration of scientific process spanning the entire tree of life, BGI-Research and collaborating institutes report three groundbreaking studies that showcase how cutting-edge CycloneSEQ nanopore sequencing technology has successfully decoded complete genomes from a bearded dragon and African wild rice to telomere-to-telomere perfection, plus circular bacterial genomes. This work highlights a new era in understanding life's blueprint across animals, plants, and microorganisms.

 Three CycloneSEQ breakthrough publications spanning animals, plants, and microorganisms.
Three CycloneSEQ breakthrough publications spanning animals, plants, and microorganisms.
By reading much longer stretches of DNA and combining them with accurate short reads, scientists have now built complete "instruction manuals" that reveal how these organisms function, evolve, and can be improved for human benefit. These studies provide high-contiguity assemblies and quantitative performance benchmarks, alongside testable hypotheses for sex determination in reptiles, resources for rice improvement, and practical guidance for closing bacterial genomes.
The complete genome reveals genes involved in reptile sex determination
In one study, researchers assembled a near-complete, chromosome-scale genome for the bearded dragon Pogona vitticeps, a lizard whose sex can be set by chromosomes or shifted by temperature during development. The team produced an almost end‒end DNA map of all 16 chromosomes and pinpointed a compact region on the Z sex chromosome where male and female differences are concentrated.
CycloneSEQ long-read sequencing combined with short-read polishing yielded a 1.79-Gb assembly with a contig N50 of 202.5 Mb, 16 fully anchored chromosomes, and 31 of 32 telomeres placed. Approximately 83% of the Z chromosome is pseudo-autosomal, whereas a small sexually differentiated region comprises distinct evolutionary strata.
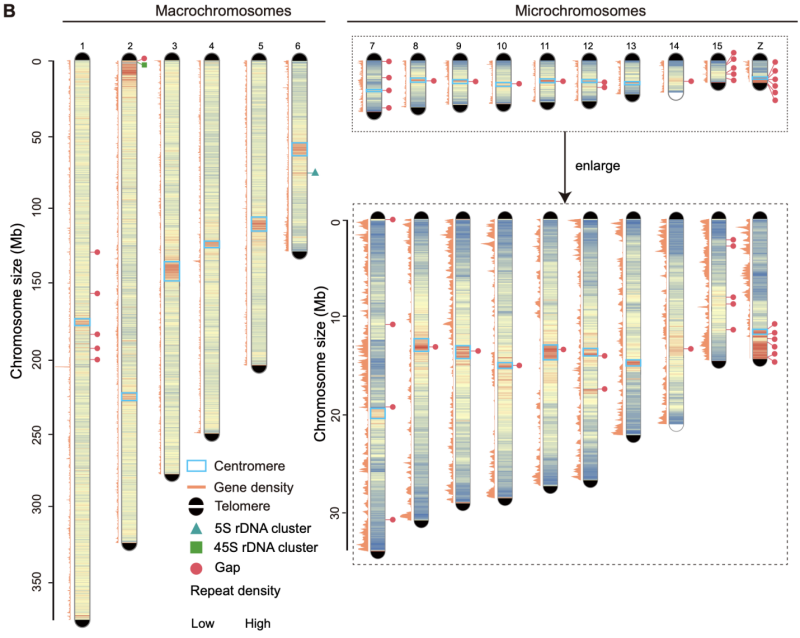 Assembly quality metrics showing 202.5 Mb contig N50 and 97.6% BUSCO completeness advance reptile genomics with a high-quality reference genome genetic and enable temperature-dependent sex determination research.
Assembly quality metrics showing 202.5 Mb contig N50 and 97.6% BUSCO completeness advance reptile genomics with a high-quality reference genome genetic and enable temperature-dependent sex determination research.
Within the SDR, the study identified AMH (anti-Müllerian hormone) together with its receptors AMHR2 and BMPR1A; AMH on the Z (AMH-Z) shows significantly male-biased expression at the earliest examined stage of gonadal differentiation and remains elevated thereafter. The authors propose, cautiously, that AMH-Z is a strong candidate for the master sex-determining gene in this species and outline a model in which a duplicated copy of autosomal AMH translocated to the proto-sex chromosome and helped launch Z/W divergence. These inferences, while supported by expression and evolutionary analyses, will benefit from future functional tests.
Gap-free genomes boosts climate-resilient rice breeding
A second study delivers a telomere-to-telomere, gap-free reference genome for African wild rice, a species valued for its perennial growth, robust biomass, and disease resistance. All 12 chromosomes are now assembled end-to-end with all telomeres and centromeres, offering an intact genetic blueprint to accelerate breeding.
Despite the challenging heterozygosity level of approximately 1.63% and high repeat content (~46.23% of the genome), researchers overcame these assembly obstacles using a hybrid approach that combined PacBio HiFi reads, Hi-C scaffolding, and CycloneSEQ ultra-long reads exceeding 79 kb in length to produce a 331 Mb, gap-free assembly covering 24 telomeres and 12 centromeres, with 98.6% BUSCO completeness indicating near-perfect quality.
Comparative genomics revealed extensive structural variation relative to cultivated rice species, including more than 87 Mb of large structural variants and a 30.2 Mb segmental duplication region containing 4179 gene pairs. The analysis further identified expanded disease resistance genes (654 NBS-LRRs) and 2,095 transcription factors, highlighting the genetic complexity and adaptive potential of this species.
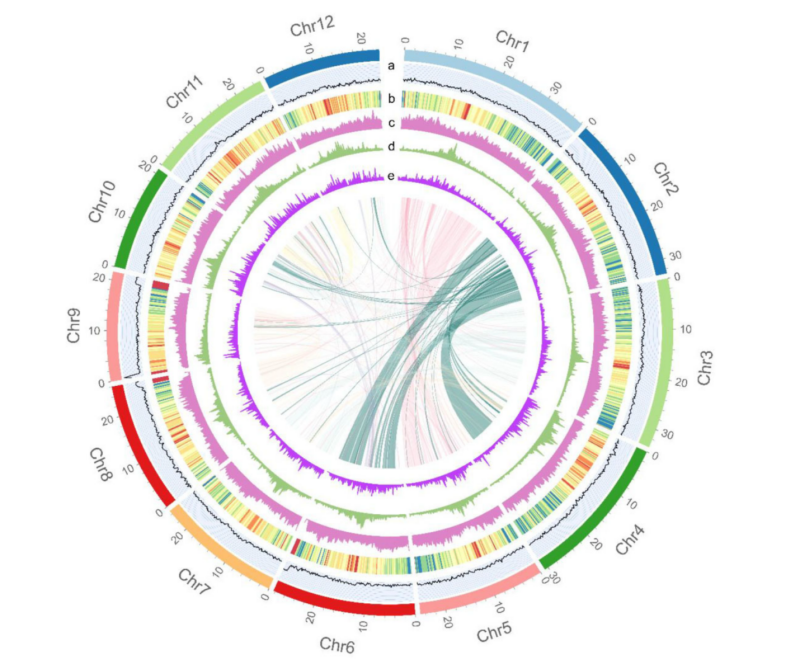 T2T genome circular plot displays a complete 12-chromosome architecture with 24 telomeres and 12 centromeres, transforms plant breeding with gap-free reference, and enables climate-resilient rice development.
T2T genome circular plot displays a complete 12-chromosome architecture with 24 telomeres and 12 centromeres, transforms plant breeding with gap-free reference, and enables climate-resilient rice development.
These results provide a high-integrity reference that can support marker development, trait mapping, and introgression of beneficial alleles into cultivated rice.
Hybrid assembly achieves complete circular genomes
A third study evaluated genome closure in bacteria via long–short hybrid assembly. Mixing long reads from CycloneSEQ with accurate short reads can turn fragmented bacterial drafts into complete circular genomes. Tests across ten strains revealed that the hybrid assembly routinely produced circular chromosomes and resolved multicopy rRNA operons and plasmids that broke short-read assemblies.
The long-read-only assemblies achieved contiguity but had substantially greater base-level errors, with approximately 28 small insertions or deletions per 100,000 DNA letters in comparisons, whereas the hybrid assemblies maintained low mismatch and indel rates, often fewer than 1–10 per 100 kb depending on the short-read depth.
Data subsampling revealed that approximately 1,000 Mb of short-read data, equivalent to sequencing a bacterial genome approximately 200--500 times, combined with 100--200 Mb of long-read data was sufficient to close most genomes with high accuracy in this cohort.
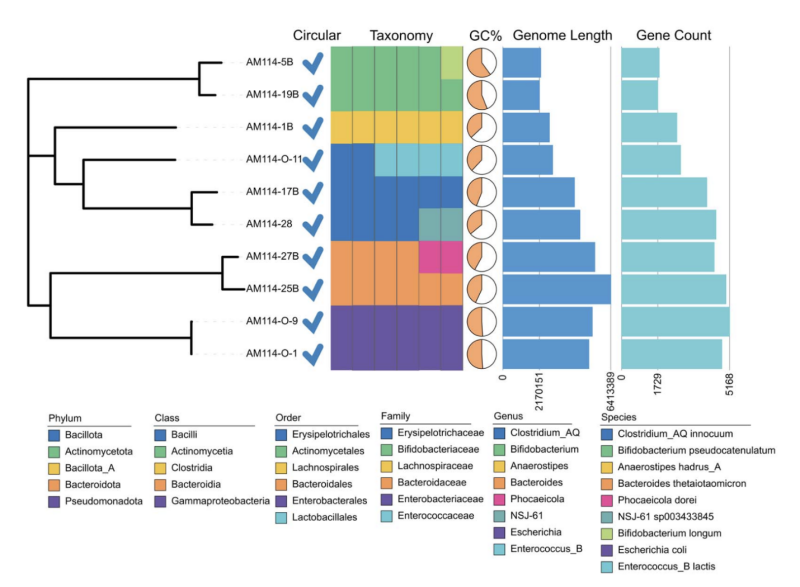 Hybrid assembly error rates of 0.08 mismatches and 0.15 indels per 100 kb versus long-read-only methods revolutionize microbiology with complete circular genomes and enable accurate pathogen analysis and antibiotic resistance studies.
Hybrid assembly error rates of 0.08 mismatches and 0.15 indels per 100 kb versus long-read-only methods revolutionize microbiology with complete circular genomes and enable accurate pathogen analysis and antibiotic resistance studies.
In metagenomic mock communities, hybrid methods yielded more single-contig, higher-quality metagenome-assembled genomes than long-read-only workflows under the study's computing conditions.
These collaborative studies demonstrate how combining complementary sequencing technologies (ultralong reads, accurate short reads, and long-range scaffolding) can produce complete, high-quality genomes across diverse life forms. This work provides valuable resources for reptile sex determination research, rice breeding programs, and microbial genomics applications worldwide.
All three studies can be accessed here:
A near-complete genome assembly of the bearded dragon Pogona vitticeps provides insights into the origin of Pogona sex chromosomes
https://academic.oup.com/gigascience/article/doi/10.1093/gigascience/giaf079/8237438
Telomere-to-telomere African wild rice (Oryza longistaminata) reference genome reveals segmental and structural variation
https://academic.oup.com/gigascience/article/doi/10.1093/gigascience/giaf074/8237436
Efficiently constructing complete genomes with CycloneSEQ to fill gaps in bacterial draft assemblies
https://gigabytejournal.com/articles/154



