Cell-free DNA (cfDNA) is made of tiny DNA fragments released into the bloodstream when cells break down. Because these fragments carry valuable biological signals, cfDNA has become a cornerstone of modern “liquid biopsy” approaches, including non-invasive prenatal testing (NIPT) and emerging tools for early cancer detection. Yet for years, scientists have faced a basic mystery: why do cfDNA fragments consistently show certain “end patterns” at their broken ends, and why do these patterns differ from person to person?
 The Study “cfGWAS reveal genetic basis of cell-free DNA end motifs” was published in Nature Communications.
The Study “cfGWAS reveal genetic basis of cell-free DNA end motifs” was published in Nature Communications.
In a recent paper published in Nature Communications, researchers from BGI-Research, working with Wuhan Children’s Hospital and multiple partner institutions, report the first genome-wide study designed to answer this question. The team conducted a genome-wide association study (GWAS) focused specifically on cfDNA end-motif frequencies, a method they call cfGWAS. Using data from 28,016 pregnant women, the study identifies inherited genetic factors that help shape cfDNA end patterns across the whole genome, and then confirms key findings using independent datasets and laboratory experiments.
A central advance of the work is that it extracts two kinds of information from the same routine clinical sequencing data. From low-depth whole-genome sequencing originally generated for NIPT, the researchers inferred genetic variation across the genome, and at the same time measured how often each of 256 possible four-letter (4-mer) DNA “end motifs” appears at the 5’ ends of cfDNA fragments. In plain terms, one set of sequencing reads was used both to learn “what genetic differences people carry” and to learn “what cfDNA end patterns their blood contains,” creating a direct bridge between genetics and cfDNA fragment signatures at population scale.
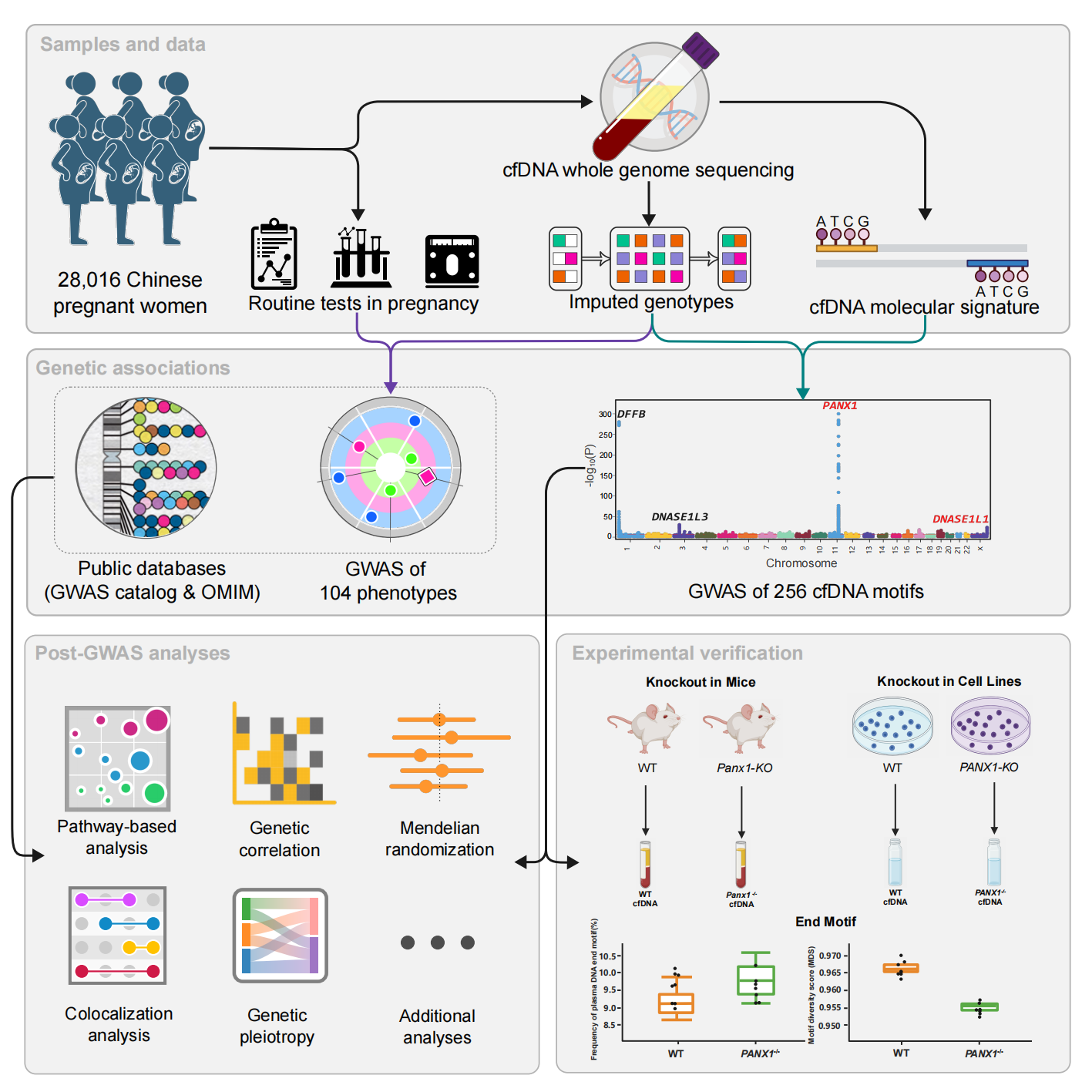 Overview of the study design.
Overview of the study design.
“This is like using a single key to open two doors at once - one to genetic information and the other to molecular phenotypes,” said Dr. Xin Jin, the project lead at BGI-Research and corresponding author of the paper. “cfGWAS is the first systematic effort to answer the fundamental question of which genetic differences determine each person’s unique cfDNA fragment patterns in the blood.”
The analysis revealed 15 highly reliable genetic regions (loci) that influence cfDNA end motifs, together affecting 176 different motif types. One of the most important discoveries involves PANX1, a gene that encodes Pannexin-1, a channel protein in the cell membrane known to participate in cell signaling, development, inflammation, and forms of cell death. PANX1 had not previously been confirmed as a regulator of cfDNA fragment end patterns, but in this genome-wide analysis it stood out as one of the strongest genetic influences on cfDNA end motifs.
Some results strongly reaffirmed what scientists suspected from earlier smaller or more targeted studies. Two classic DNA-cutting enzymes (nucleases) involved in DNA fragmentation and clearance, DFFB and DNASE1L3, emerged again as major players, providing large-population confirmation that these enzymes help shape the fragment patterns seen in blood. The study also highlighted DNASE1L1, a related gene whose connection to cfDNA biology had not been firmly established in living systems.
To make sure the findings were not specific to one dataset, the team tested them in three independent replication resources, including an additional large Chinese NIPT cohort, a Dutch NIPT genetics resource, and a smaller general-population cohort. Most of the strongest signals seen in the discovery cohort were also observed again, supporting the reliability of the genetic links.
The researchers then moved from statistics to biology by testing PANX1 directly in the lab. They created Panx1 knockout mice and also generated PANX1 knockout cell lines. The mouse experiment demonstrates that when PANX1 was missing, cfDNA end-motif patterns shifted measurably, and cfDNA concentration tended to rise. These results provide functional evidence that PANX1 can influence how cfDNA is produced, processed, and/or cleared from the bloodstream.
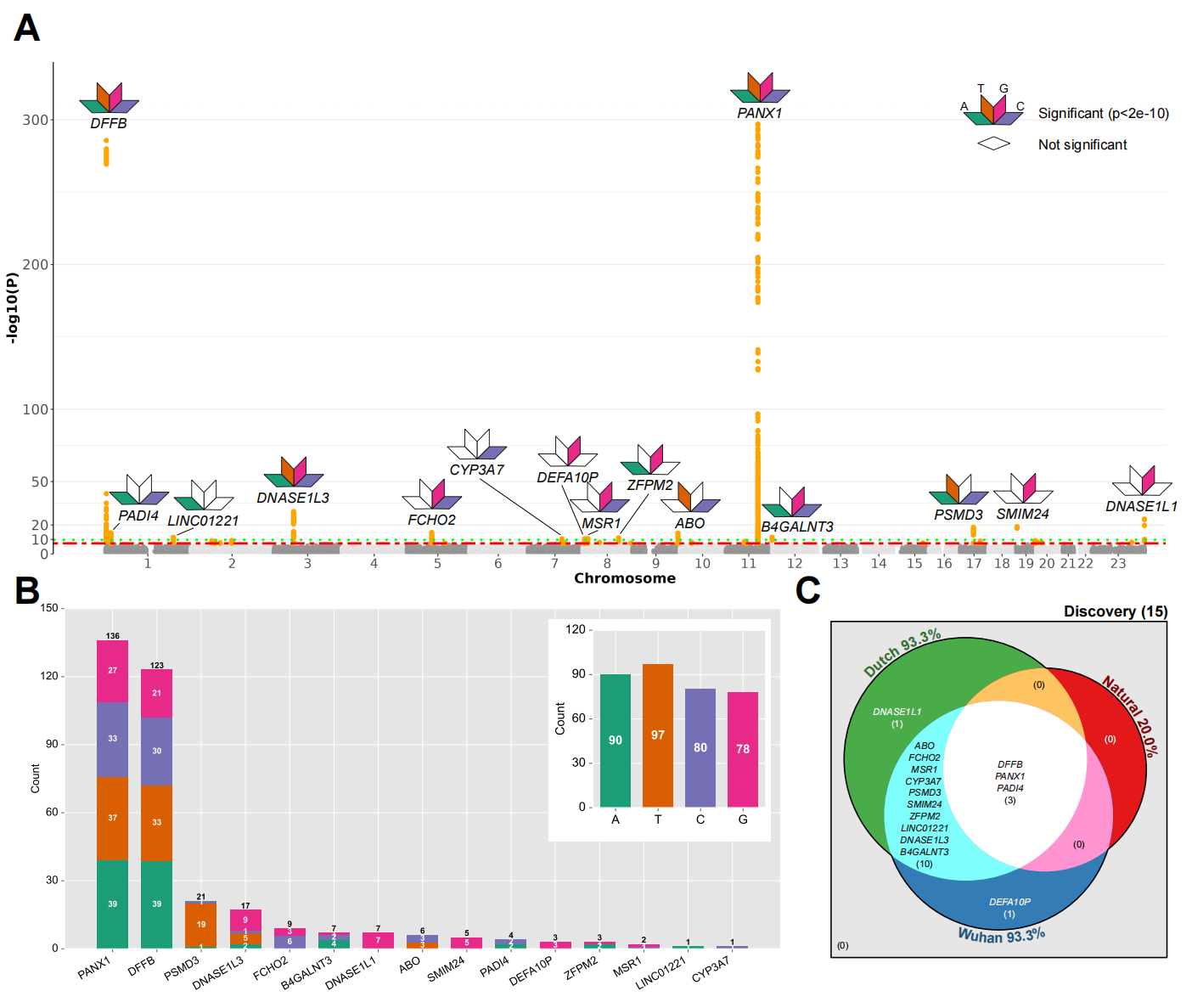 The results for the GWAS analysis of 256 motifs.
The results for the GWAS analysis of 256 motifs.
The study also looked beyond individual genes to ask what kinds of cells might be driving these cfDNA signatures. Using genetic causal inference methods, the researchers found strong evidence that white blood cells - particularly neutrophils - have a causal impact on cfDNA end-motif features. This supports a biologically intuitive picture: immune cells are both an important source of circulating cfDNA and a key factor shaping how cfDNA looks in the blood, helping explain why cfDNA profiles can change in conditions linked to inflammation and immune activity.
The practical significance is clear for the future of liquid biopsy. Because people’s inherited genetics can affect cfDNA fragmentation and clearance, understanding these “genetic background” effects could help improve the accuracy and consistency of cfDNA-based tests. This may be especially valuable in early cancer detection, where tumor-derived DNA can be extremely scarce and where small biases in measurement may matter. The discovery of new regulators such as PANX1 also suggests fresh directions for exploring drug targets in diseases where cfDNA clearance is abnormal, including certain autoimmune and inflammatory conditions. In pregnancy-related medicine, clarifying the genetic basis of cfDNA fragment signatures may support more stable and interpretable models for risk prediction when cfDNA patterns are used as biomarkers.
Globally, cfDNA sequencing has already been performed at massive scale through NIPT and other clinical testing, producing a large but underused resource of genomic data. This study demonstrates that these existing datasets can be re-analyzed to uncover fundamental human biology at genome-wide scale. As sample sizes grow, cfGWAS may reveal additional genes and pathways that govern cfDNA dynamics, accelerating both basic research and next-generation liquid biopsy applications.
Each participant provided informed consent before enrollment. Ethics approval for this study is obtained.
This study can be accessed here: https://www.nature.com/articles/s41467-025-67940-w



