On February 18, 2026, researchers from BGI-Research's State Key Laboratory of Genome and Multi-omics Technologies, working with collaborators at The Chinese University of Hong Kong, Shenzhen, reported in Chemical Science a quantum–classical hybrid approach for modeling electron transfer in a biological protein environment. The open-access study introduces a novel framework called VQE-PDFT, which overcomes long-standing challenges in accurately simulating electron transfer in strongly correlated systems, a critical aspect in fields like biochemistry and pharmacology.
 This study “Quantum-classical hybrid computation of electron transfer in a cryptochrome protein via VQE-PDFT and multiscale modeling” was published open access in Chemical Science, demonstrating the practical application of a quantum-classical hybrid approach for calculating biological electron transfer rates.
This study “Quantum-classical hybrid computation of electron transfer in a cryptochrome protein via VQE-PDFT and multiscale modeling” was published open access in Chemical Science, demonstrating the practical application of a quantum-classical hybrid approach for calculating biological electron transfer rates.

Outside back cover feature. By highlighting the study in the journal's cover feature, this artwork signals editorial visibility while pointing readers to the underlying quantum–classical electron-transfer result.
A long-standing barrier in accurate electron-transfer simulation is the inherent strongly correlated system, where multiple electronic configurations and instantaneous electron–electron interactions both matter but are difficult to capture efficiently with conventional methods. In VQE-PDFT, the quantum circuit handles the most quantum-mechanically demanding part of the calculation and measures the resulting density matrices, while a classical computation step recovers much of the remaining correlation energy. This division of labor keeps the quantum component shallow and feasible, while still supporting chemically meaningful energies.
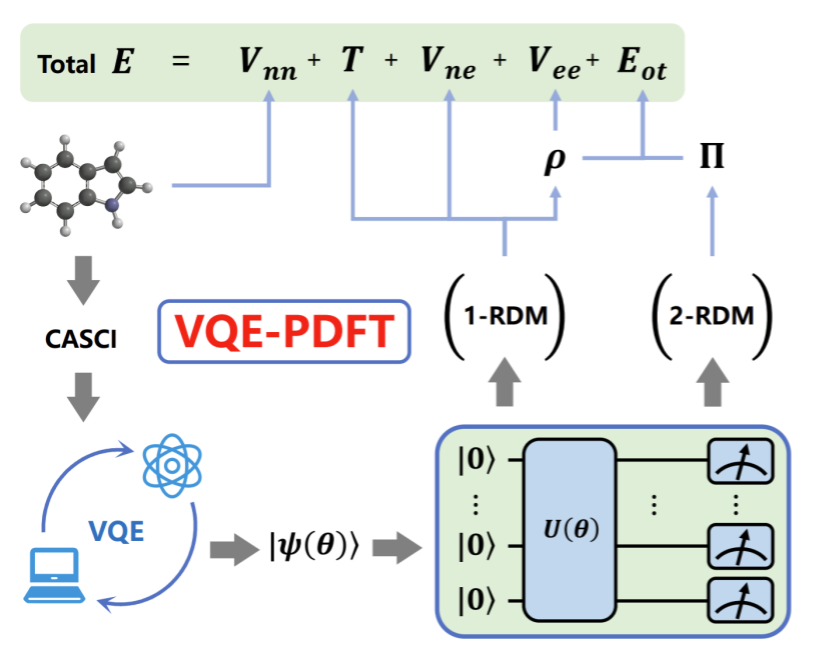
VQE-PDFT workflow. By using a quantum circuit to obtain reduced density matrices and a classical PDFT step to recover correlation energy, the framework turns near-term quantum outputs into chemically usable energetics.
To demonstrate the method in a biologically relevant setting, the team embedded VQE-PDFT into a multiscale architecture and focused on electron transfer between adjacent tryptophan residues inside the European robin cryptochrome protein ErCRY4. Cryptochromes are widely studied in connection with hypotheses about avian magnetoreception; here, the protein serves as a stringent test case for computing an electron-transfer step in a realistic electrostatic environment, without claiming to resolve bird navigation mechanisms on its own.
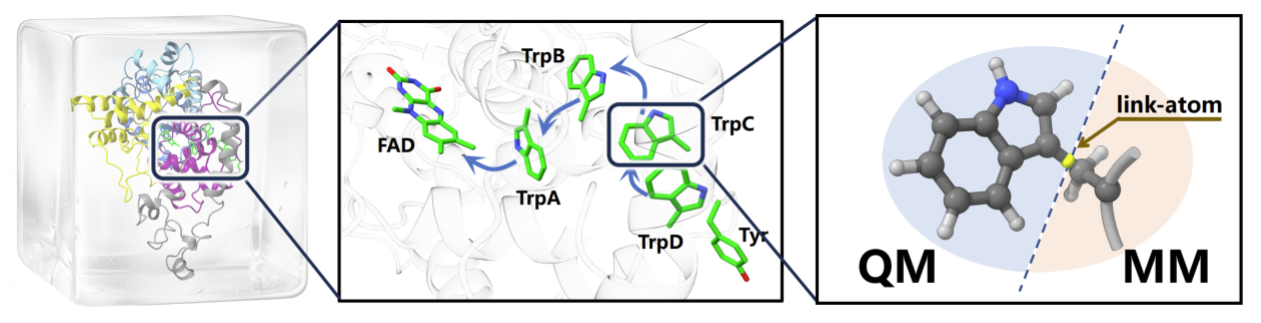
QM/MM embedding of the electron-transfer region. By isolating a tryptophan-centered quantum region within a classical protein environment, the model evaluates transfer-driving energetics under biologically relevant electrostatics.
A key engineering step was designing compact quantum circuits tailored to the two electronic states involved in the transfer. The paper reports shallow circuit depths of 4 and 6 layers for the two circuit types, compared with 35 layers for a deeper reference circuit, illustrating how system-specific circuit design can make calculations feasible on today's noisy hardware without sacrificing accuracy for the chosen active spaces.
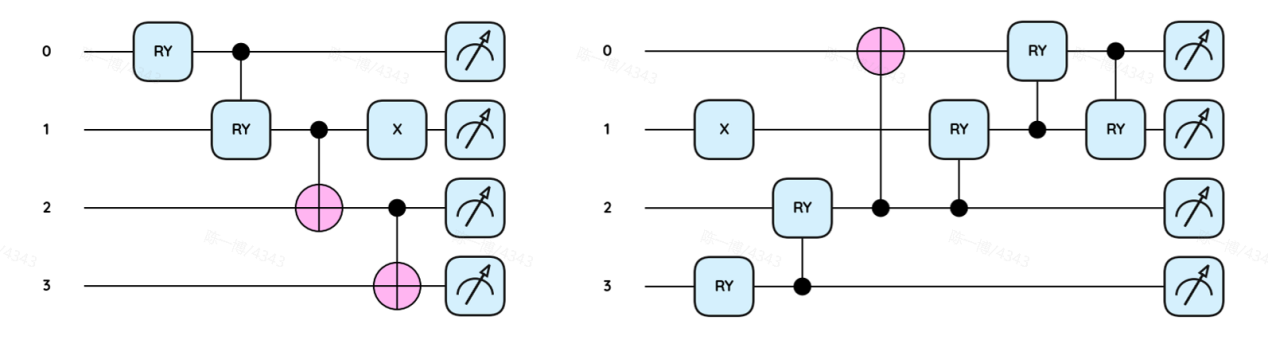 Shallow, state-specific circuits. By tailoring compact ansatz circuits for closed- and open-shell cases, the study reduces circuit depth to fit NISQ constraints while retaining expressibility for the chosen active spaces.
Shallow, state-specific circuits. By tailoring compact ansatz circuits for closed- and open-shell cases, the study reduces circuit depth to fit NISQ constraints while retaining expressibility for the chosen active spaces.
Across 20 protein conformations, the electron-transfer rate was estimated to be on the order of 10¹⁰ s⁻¹, close to the ~0.7 × 1010 s⁻¹ value reported by ultrafast spectroscopy, while conventional DFT calculation yields transfer rate on the order of 1011 s-1. The authors present this agreement as evidence that a carefully constrained quantum–classical workflow can reach experimentally relevant kinetics for this protein electron-transfer step, while acknowledging the deliberate simplifications inherent in compact active-space modeling.
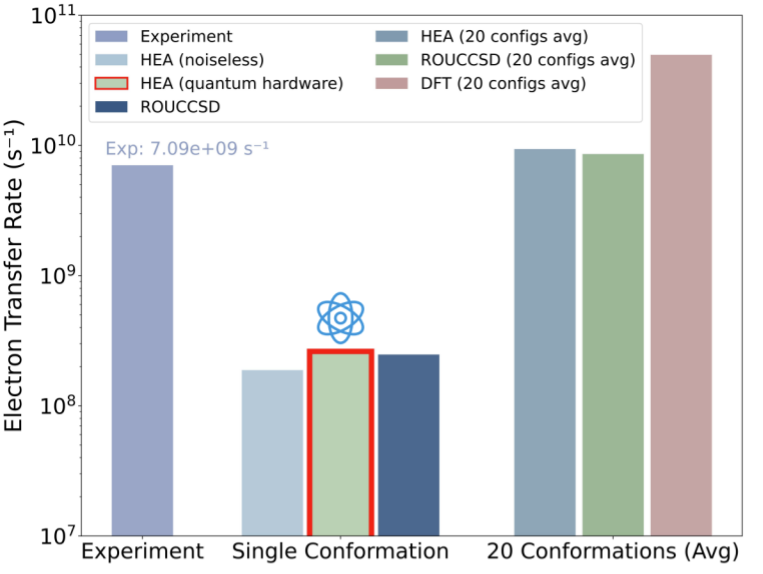
Transfer-rate benchmarking across conformations. By comparing predicted rates against ultrafast spectroscopy and classical baselines, the plot shows the hybrid workflow reaching the correct order of magnitude for this cryptochrome transfer step.
As a proof-of-concept hardware validation, the team executed density-matrix measurements for one ErCRY4 conformation on a customized 13-qubit superconducting device and analyzed how noise affects both single-point energies and derived transfer-rate parameters. The paper reports that hardware noise can introduce systematic shifts in absolute energies, but that computing energy differences to build those parameters can partially cancel the shifts, preserving usable estimates under current hardware limits. The authors treat this as an early demonstration and call for broader validation as hardware and software workflows mature.
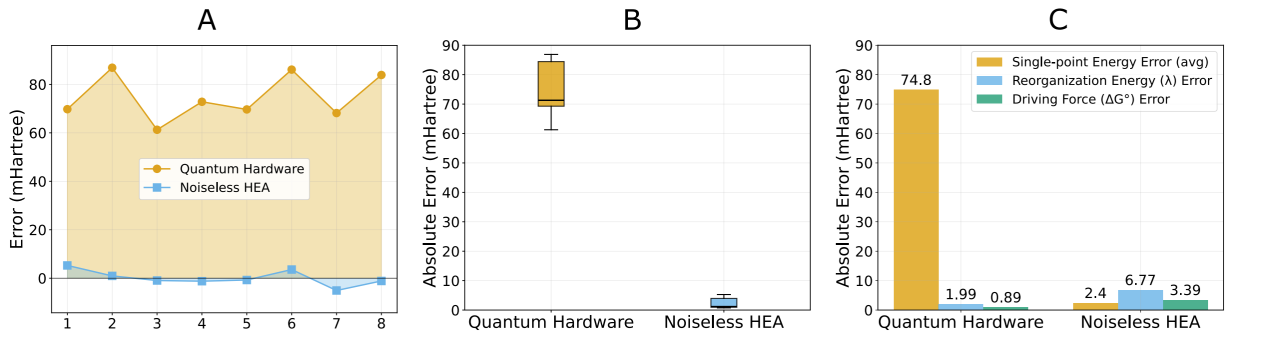 Noise propagation and partial cancellation. By separating single-point energy errors from Marcus-parameter errors, the analysis illustrates why difference-based observables can be more robust than absolute energies on noisy hardware.
Noise propagation and partial cancellation. By separating single-point energy errors from Marcus-parameter errors, the analysis illustrates why difference-based observables can be more robust than absolute energies on noisy hardware.
Taken together, this study carries three broader messages. First, quantum computers do not need to replace classical computers to be useful in biology. They can act as a specialist called in for the most intensive part of a calculation, passing results back to classical computation that handle the rest. This kind of targeted partnership, demonstrated here for a protein electron-transfer step, offers a realistic near-term path for quantum hardware to contribute to drug discovery, enzyme design, and other areas where the quantum behavior of electrons in molecules matters. Second, even imperfect quantum hardware can give useful answers when the question is framed as a relative comparison. Systematic noise that shifts all measurements in the same direction tends to partially cancel when the result of interest is a difference, much as a slightly out-of-tune instrument can still play a recognizable melody if all notes are shifted equally. Third, the computational framework developed here is designed to grow with the technology. As quantum processors become more powerful, the same approach can be applied to a larger active space and more complex biological processes without rebuilding the pipeline from scratch, making this work a concrete early step toward a future where quantum computing routinely supports biomedical research.
This research can be accessed at https://doi.org/10.1039/D5SC07528A.



