The human gut harbors vast numbers of bacteriophages that help shape microbial communities, yet for most of these phages the bacterial host remains unknown. In the Gut Phage Database (GPD), hosts had been assigned to fewer than 29% of 142,809 cataloged phage genomes, leaving much of the gut virome as biological "dark matter" and limiting efforts to link phage-bacteria interactions to conditions such as obesity and inflammatory bowel disease.
 A dual-track strategy for exploring bacteriophage dark matter integrates a large-scale experimental collection as the upper track and ai prediction as the lower track.
A dual-track strategy for exploring bacteriophage dark matter integrates a large-scale experimental collection as the upper track and ai prediction as the lower track.
To address this challenge, researchers from BGI-Research and partner institutes published two complementary studies in Nature Communications: one establishes the Gut Phage Biobank (GPB), a large-scale experimental collection of cultivated phage isolates, while the other deploys the AI-driven host-prediction framework VirHost Hunter to decode viral dark matter at scale.
Experimental Track: Building the Gut Phage Biobank
On the experimental track, the GPB study consolidated classical plaque assays, repetitive enrichment co-culture, and metagenomics-guided screening into a systematic workflow that yielded 104 phage isolates targeting 29 gut bacterial species selected for high abundance or disease association.
 The companion Gut Phage Biobank study provides the experimental counterpart to VirHost Hunter, together forming a dual-track strategy for decoding gut phage "dark matter," published in Nature Communications (2025).
The companion Gut Phage Biobank study provides the experimental counterpart to VirHost Hunter, together forming a dual-track strategy for decoding gut phage "dark matter," published in Nature Communications (2025).
The three methods proved complementary: the classic approach yielded the most isolates, repetitive enrichment was essential for high-diversity samples, and metagenomics-guided screening enabled precise targeting of obligate anaerobes.
Genomic analysis placed 39 isolates into clusters with no RefSeq representatives and identified an undescribed viral family and four undescribed genera among phages infecting obligate anaerobes.
Two Mediterraneibacter gnavus phages of an uncharacterized genus showed global prevalence exceeding that of the well-known p-crAssphage, suggesting the current picture of dominant gut phage lineages may be incomplete.
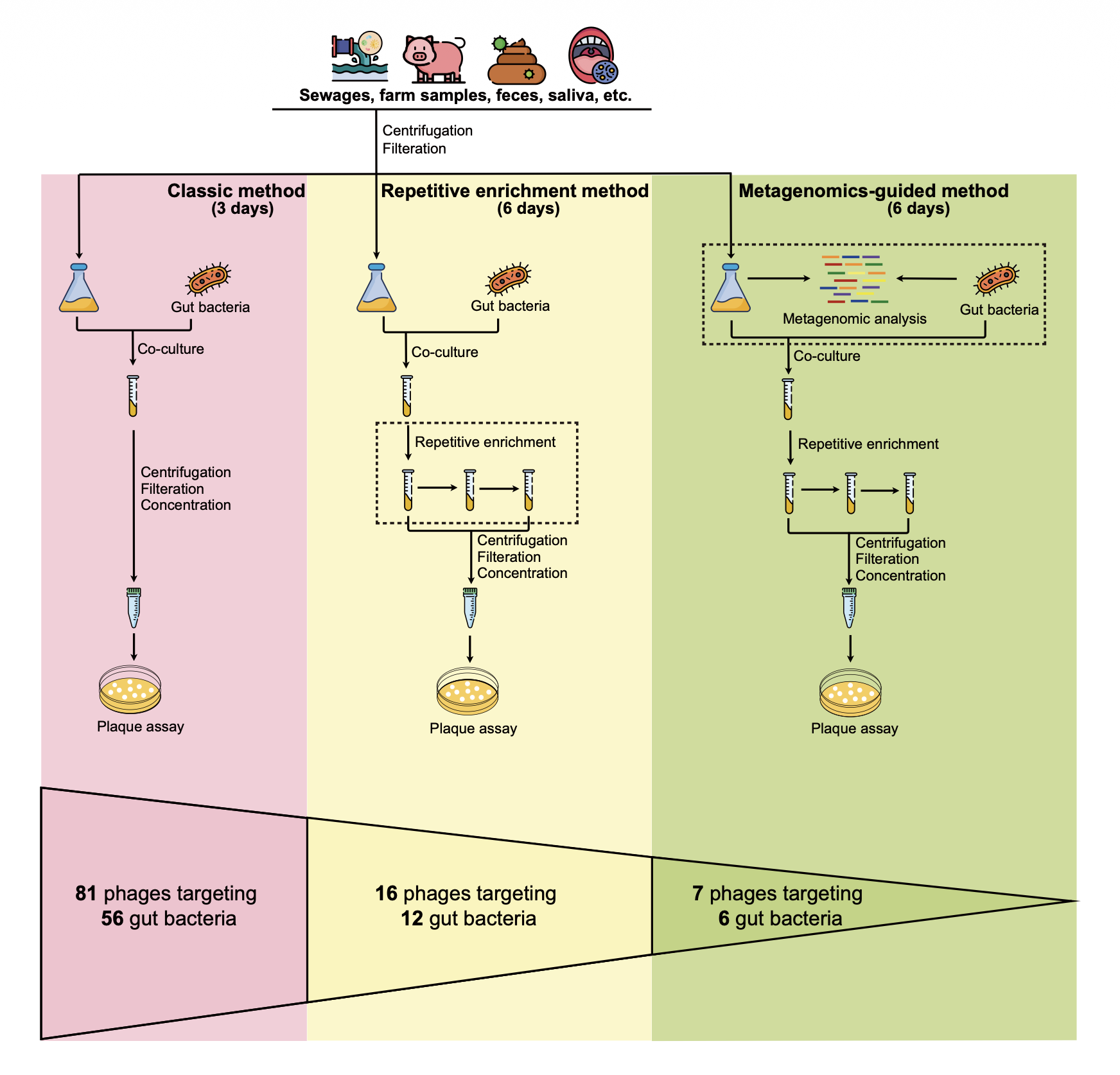 Three complementary phage isolation methods, classical, repetitive enrichment, and metagenomics-guided, capture different ecological niches and maximize the recovery of cultivable gut phages.
Three complementary phage isolation methods, classical, repetitive enrichment, and metagenomics-guided, capture different ecological niches and maximize the recovery of cultivable gut phages.
Cross-infection experiments under both oxic and anoxic conditions showed that GPB phages exhibit high host specificity at the species and strain level, with oxygen availability influencing both host range and bactericidal activity. In vivo, phage CPB1092 targeting Dorea longicatena, a gut bacterium implicated in both obesity and type 2 diabetes, significantly reduced the relative abundance of its target in a mouse model, providing early evidence that cultivated phages can selectively modulate specific members of the gut microbiota.
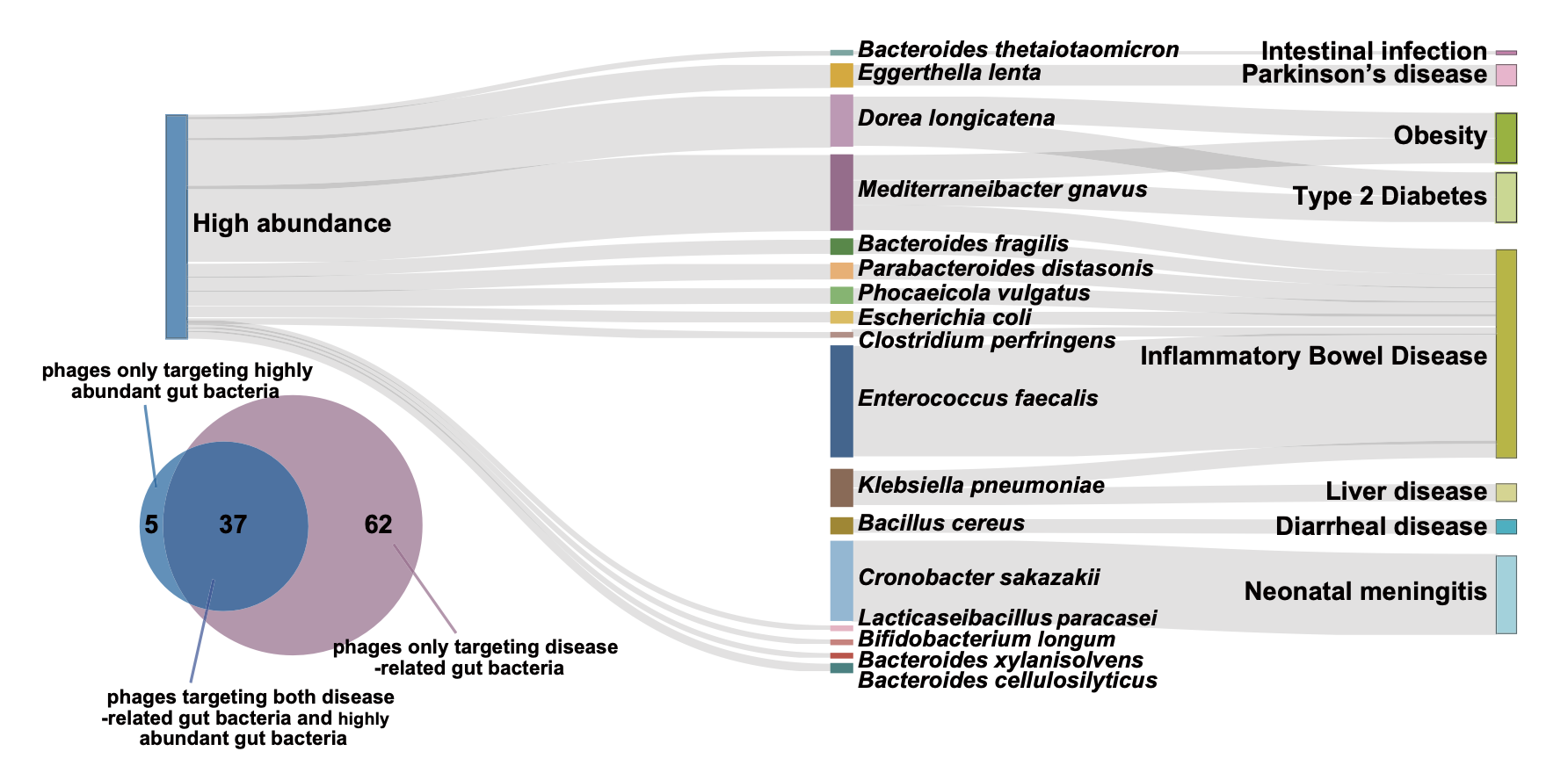 Host-species mapping of GPB phages to disease-associated gut bacteria illustrates the biobank's focus on clinically relevant targets across conditions including obesity, diabetes, and inflammatory bowel disease.
Host-species mapping of GPB phages to disease-associated gut bacteria illustrates the biobank's focus on clinically relevant targets across conditions including obesity, diabetes, and inflammatory bowel disease.
Computational Track: Developing the VirHost Hunter AI Framework
On the computational track, BGI-Research and Shenzhen University developed an AI framework named “VirHost Hunter”, which focuses on tail proteins and lysins, combining protein-language-model representations from ProtT5 with DNA-sequence features captured through a Vision Transformer to enable host prediction even from incomplete assemblies.
 VirHost Hunter, an AI-driven framework for high-resolution phage-host assignment using key proteins and large language models, published in Nature Communications (2026).
VirHost Hunter, an AI-driven framework for high-resolution phage-host assignment using key proteins and large language models, published in Nature Communications (2026).
In a validation set of 156 cultivated gut phages, VirHost Hunter at a 95% precision cutoff correctly identified hosts for 73 phages at the family level and 58 at the genus level, whereas the CRISPR-based method yielded no assignments at that threshold.
Combining the two approaches raised the GPD host assignment ratio from 28.66% to 62.66%, covering 58 species including 42 obligate anaerobes, and revealing phages predicted to target Akkermansia muciniphila and Prevotella copri.
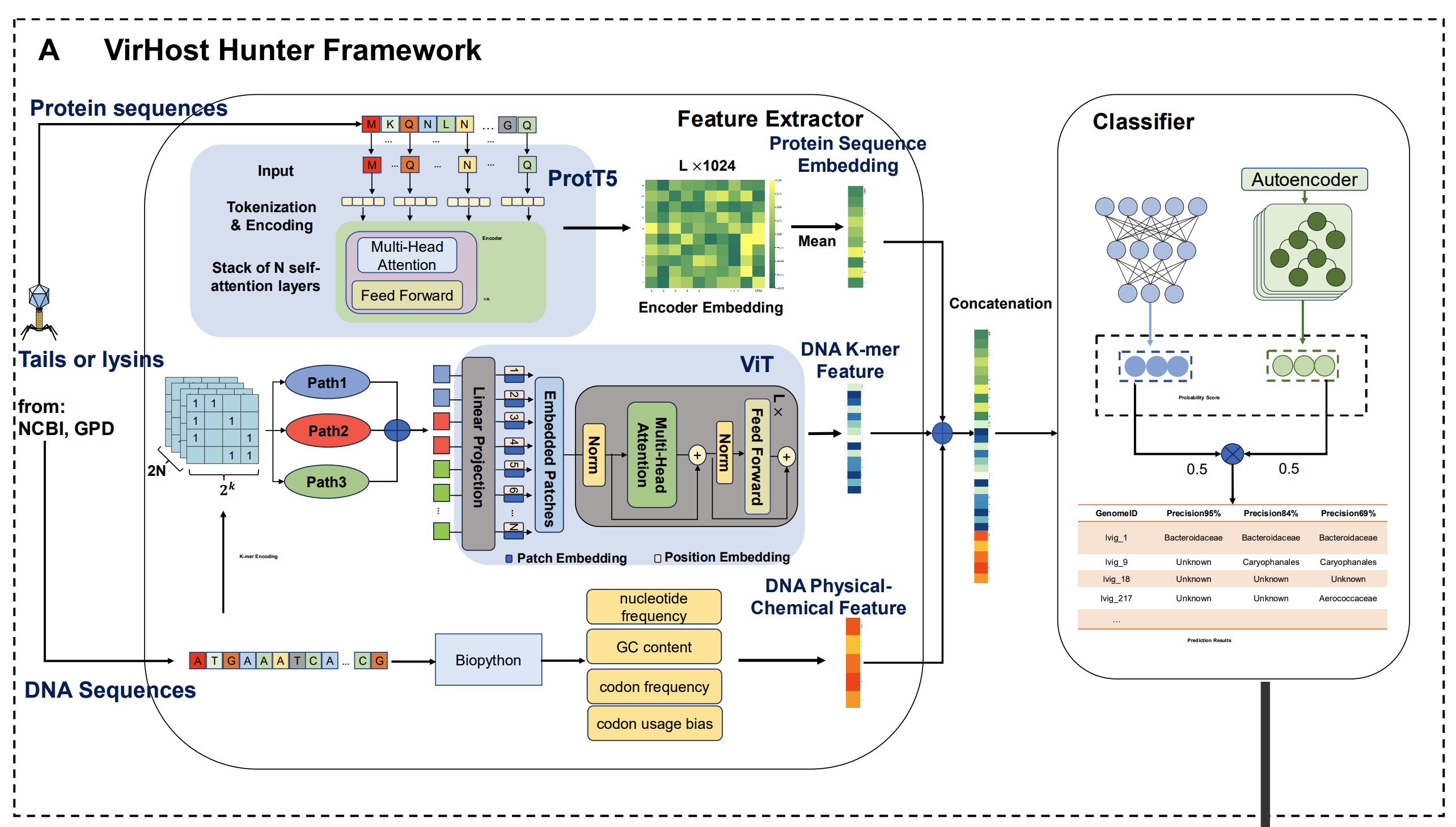 The VirHost Hunter framework integrates ProtT5-based protein embeddings with Vision Transformer DNA-sequence features, enabling host prediction from tail proteins and lysins even when full genomes are unavailable.
The VirHost Hunter framework integrates ProtT5-based protein embeddings with Vision Transformer DNA-sequence features, enabling host prediction from tail proteins and lysins even when full genomes are unavailable.
Building on these predictions, the team established the Gut Phage Lysin Database (GPLD) containing 117,698 lysins targeting 29 disease-related gut bacterial families. As a proof of concept, a synthesized lysin called Ply491_6 reduced the turbidity of Megamonas rupellensis, an obesity-associated gut bacterium, at concentrations as low as 20 µg/mL within 150 minutes while showing minimal impact on other gut bacteria tested. Further engineering would be needed before downstream application.
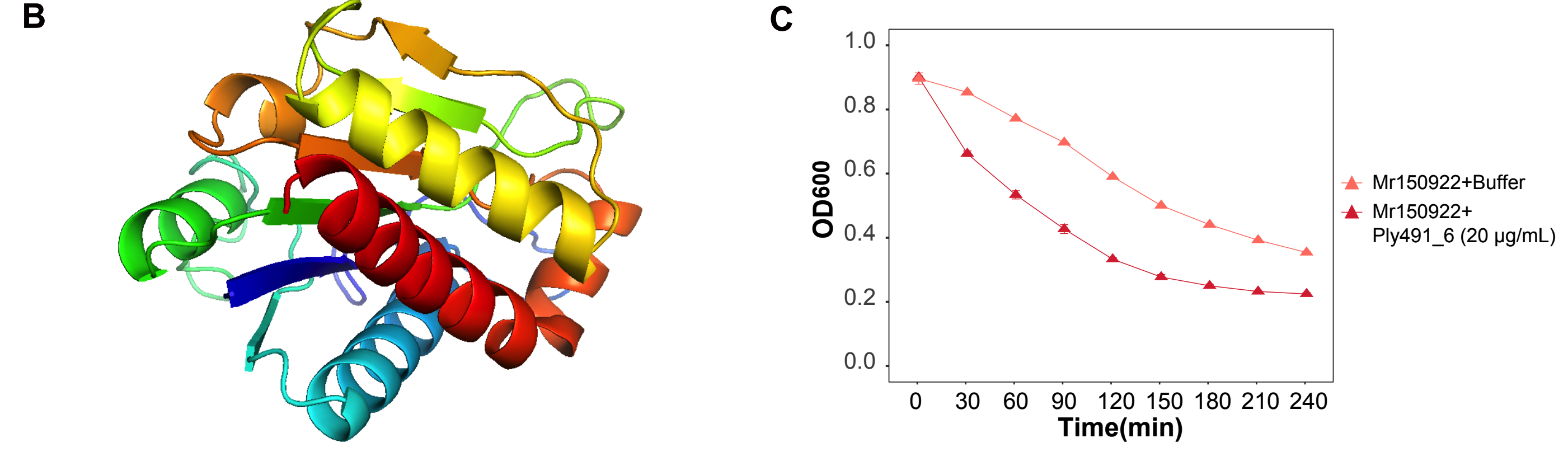 Predicted structure and in vitro lytic activity of Ply491_6 against Megamonas rupellensis illustrate how computational mining of the GPLD can yield experimentally testable candidates for selective bacterial inhibition.
Predicted structure and in vitro lytic activity of Ply491_6 against Megamonas rupellensis illustrate how computational mining of the GPLD can yield experimentally testable candidates for selective bacterial inhibition.
Together, the two studies form a replicable paradigm in which cultivated phages provide ground-truth validation for AI models while VirHost Hunter extends host prediction to uncultivable sequences, with outputs such as the GPLD feeding back into experimental pipelines.
All bacterial strains used for lysin testing were isolated from human fecal samples, and the study was conducted in accordance with applicable ethical and regulatory requirements.
The Gut Phage Biobank study is available at https://doi.org/10.1038/s41467-025-61946-0, and the VirHost Hunter study at https://doi.org/10.1038/s41467-026-70613-x.



