Wheat is a staple food for more than one-third of the world’s population, and improving its yield and quality is essential for global food security. On April 1, the State Key Laboratory of Genome and Multi-omics Technologies, initiated by BGI-Research, together with scientists from 34 universities and research institutes across 9 countries, including Xianghu Laboratory, The University of Adelaide, and Murdoch University, published a forward-looking perspective in the internationally renowned journal Nature Genetics, titled “The potential of wheat spatial omics”.
The article systematically outlines the application potential and future directions of spatial omics in wheat research, providing a new research paradigm for dissecting complex agronomic traits and breeding high-yield, high-quality wheat varieties adapted to climate change.
This marks another important achievement of the Wheat Spatial Omics Consortium (WSOC) since its establishment in November 2025.
 The Perspective Article “The potential of wheat spatial omics” was published in Nature Genetics.
The Perspective Article “The potential of wheat spatial omics” was published in Nature Genetics.
Spatial omics technologies preserve tissue architecture while revealing the spatial and temporal features of gene expression, protein distribution and metabolite changes. By overcoming the limitations of conventional omics approaches, which often lack precise cellular and tissue localization, spatial omics has become a transformative tool in life science research.
These technologies have already been widely applied in medical and animal sciences, and are beginning to show promise in plant research, including studies in rice, onion and other species. Given the complexity of the wheat genome, the multiple challenges posed by climate change, and the urgent need for precision breeding, spatial omics is emerging as a key breakthrough direction in wheat science.
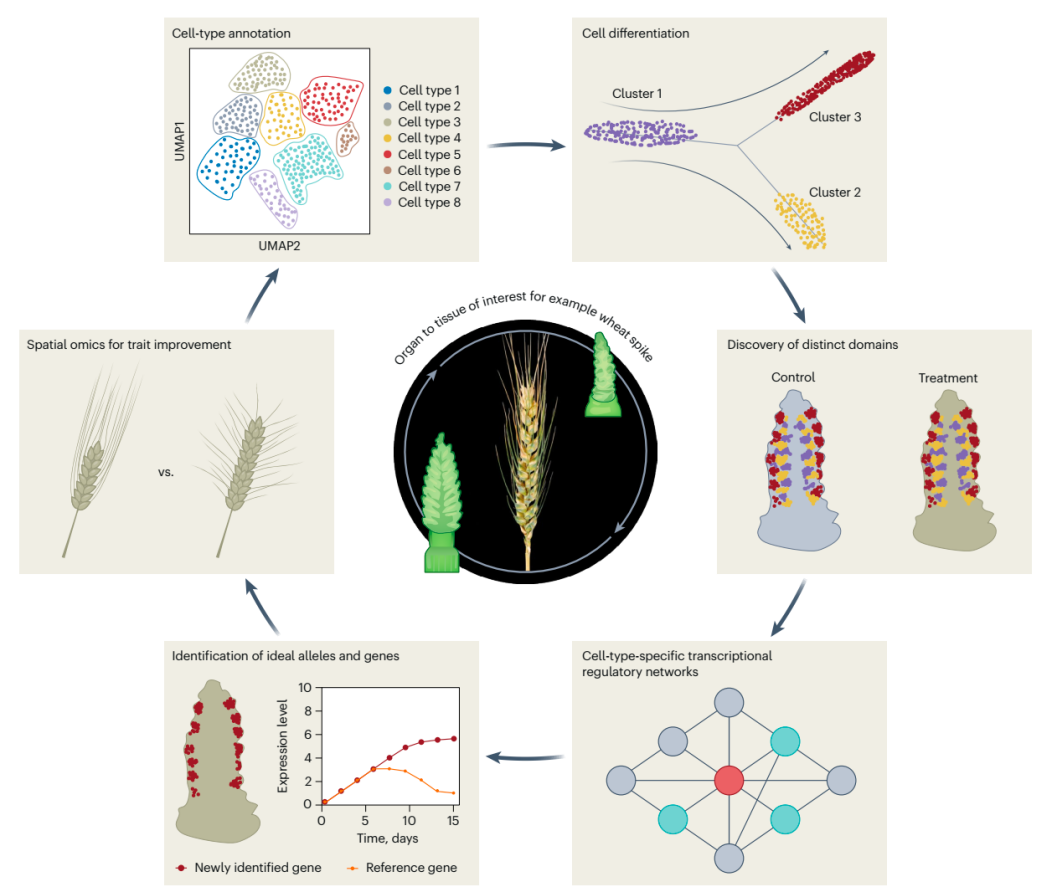 Potential applications of spatial transcriptomics in wheat.
Potential applications of spatial transcriptomics in wheat.
The perspective comprehensively reviews recent progress in spatial omics applications in plants and proposes a full experimental design framework for addressing core questions in wheat growth, development and stress response.
The authors highlight that spatial omics can generate high-resolution cellular maps of key wheat organs such as the spike, grain, root and flag leaf, helping researchers uncover molecular regulatory networks underlying spike development, grain formation, photosynthesis and nutrient uptake. It also offers a new way to understand the cell-type-specific mechanisms of wheat responses to abiotic and biotic stresses such as drought, salinity and disease.
For example, in drought research, spatial omics could allow scientists to pinpoint how specialized cells in the flag leaf - such as guard cells and subsidiary cells - change their gene expression under water deficit, thereby identifying new genes involved in stomatal regulation and providing precise targets for drought-tolerant breeding.
To address the scientific challenges posed by wheat’s highly complex genome, the paper proposes several strategies:
- Integrating multi-omics datasets: combining the wheat telomere-to-telomere reference genome, pangenome and population genomics resources to build a spatially resolved single-cell atlas covering the wheat life cycle;
- Linking datasets across scales: integrating spatial omics with genome-wide association studies (GWAS) and expression quantitative trait loci (eQTLs) to establish precise connections among genotype, phenotype and spatial gene expression;
- Technology convergence: combining single-cell multi-omics with spatial omics to better understand subgenome divergence and cell-type-specific regulatory networks, helping overcome bottlenecks in traditional gene discovery.
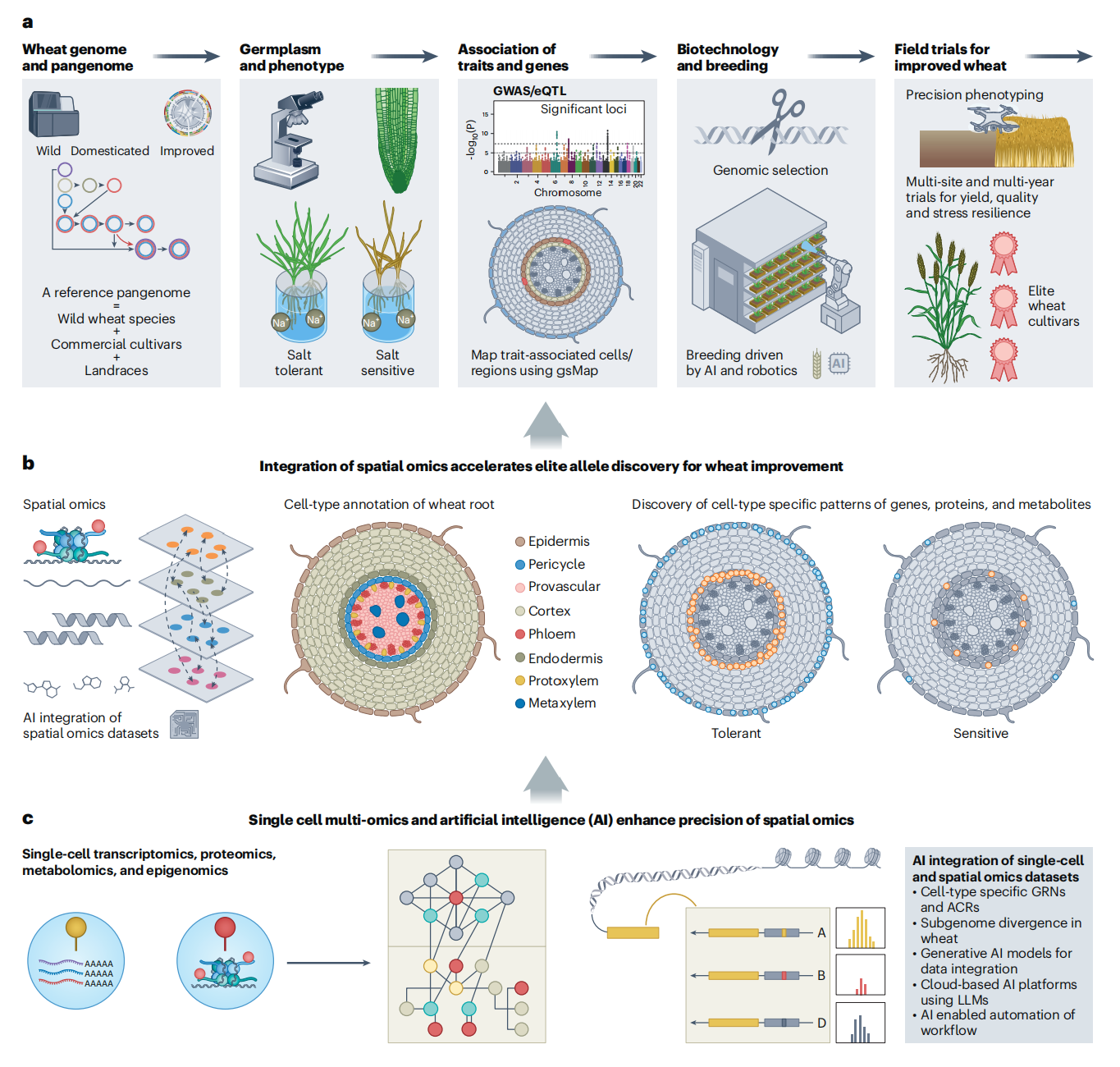
Integration of multi-omics data to dissect complex traits related to wheat genetic improvement.
Integrating multi-omics datasets: combining the wheat telomere-to-telomere reference genome, pangenome and population genomics resources to build a spatially resolved single-cell atlas covering the wheat life cycle;
Technology convergence: combining single-cell multi-omics with spatial omics to better understand subgenome divergence and cell-type-specific regulatory networks, helping overcome bottlenecks in traditional gene discovery.
The paper also systematically analyzes current technical challenges facing wheat spatial omics research and outlines possible solutions, including adopting super-resolution imaging strategies from medicine, developing plant-specific spatial omics analysis tools, and using artificial intelligence (AI) for deep analysis and modeling of spatial omics data.
To promote global collaboration in this emerging field, the research team established the Wheat Spatial Omics Consortium (WSOC) in 2025. The consortium is designed to integrate international wheat research resources, support cross-disciplinary and cross-regional collaboration, tackle technological bottlenecks in wheat spatial omics, and accelerate the translation of research discoveries into breeding applications.
The Perspective emphasizes that the integration of spatial omics with cutting-edge technologies such as genome editing, AI and precision phenotyping will help usher wheat breeding into a new era of precision design. Scientists may use AI to integrate multidimensional spatial omics data, simulate crossing and selection processes, and improve breeding efficiency. They may also discover beneficial genes from wild relatives of wheat and use them in breeding, potentially shortening breeding cycles and increasing efficiency.
This article can be accessed here: https://doi.org/10.1038/s41588-026-02542-w
About the Wheat Spatial Omics Consortium
The Wheat Spatial Omics Consortium (WSOC) was launched during the Second International Plant Spatiotemporal Omics Research Conference (STOC Plant 2025) in November 2025. As the first globally open international collaboration platform under STOC Plant dedicated specifically to wheat spatial omics research, WSOC currently brings together more than 20 research institutions and over 30 core researchers worldwide.
The consortium focuses on integrating global strengths to support large-scale wheat research, promote standardized generation of multidimensional omics data across the full wheat life cycle, construct systematic reference spatial cell atlases, and develop large AI-enabled breeding models for wheat. Its goal is to drive wheat research from static genomics toward dynamic, cell-level functional understanding, opening a critical pathway for using multi-omics big data to guide precision wheat breeding and contribute to global food security and sustainable agriculture.



