On April 16, a multi-institutional research team led by BGI-Research, the Center for Excellence in Brain Science and Intelligence Technology (CEBSIT) at the Chinese Academy of Sciences, Monash University in Australia, and other collaborators, reports in Science the discovery of an opposing molecular gradient axis as a fundamental organizing principle of the primate cerebral cortex.
By integrating spatial transcriptomics, single-nucleus RNA sequencing, magnetic resonance imaging, and neuronal tracing in the common marmoset, the team constructed a three-dimensional multimodal brain atlas and identified two molecular gradients that reconcile decades of conflicting hypotheses about how the cortex expanded during evolution. Although this is basic research, it provides a new reference map for understanding how healthy cortical organization emerges, with potential value for future studies of brain development, cognition, and disorders in which cortical architecture is altered.
The study “An opposing molecular gradient axis underlies primate cortical organization” was published in Science.
Every time we glance at a face across the room, pick out a melody from background noise, or drift into quiet reflection, distinct regions of the cerebral cortex are at work, regions that evolved and expanded over millions of years of primate history. How this remarkable outer layer of the brain came to be organized has remained one of neuroscience's most contested questions. One camp argued that the cortex grew outward from evolutionarily ancient regions involved in memory and smell; the other maintained that primary sensory areas, those governing seeing, hearing, and touch, were the original anchors. Resolving the debate required an approach that simultaneously examines gene expression, cell types, anatomy, and connectivity across the entire brain.
The team chose the common marmoset (Callithrix jacchus) as their primary model. Unlike the macaque or human brain, the marmoset's smooth, compact cortex makes whole-brain mapping tractable at single-cell resolution while retaining the essential architectural features shared across primates. Using the Stereo-seq spatial transcriptomics platform and DNBelab C4 single-nucleus RNA sequencing, with sequencing performed on DNBSEQ platforms, the team profiled nearly 500,000 nuclei from 116 brain areas and generated spatial data from 152 brain sections encompassing over 21 million segmented cells. These molecular data were coregistered with 3D MRI templates and retrograde neuronal tracing to create an integrated atlas bridging gene expression, cell composition, structure, and connectivity.
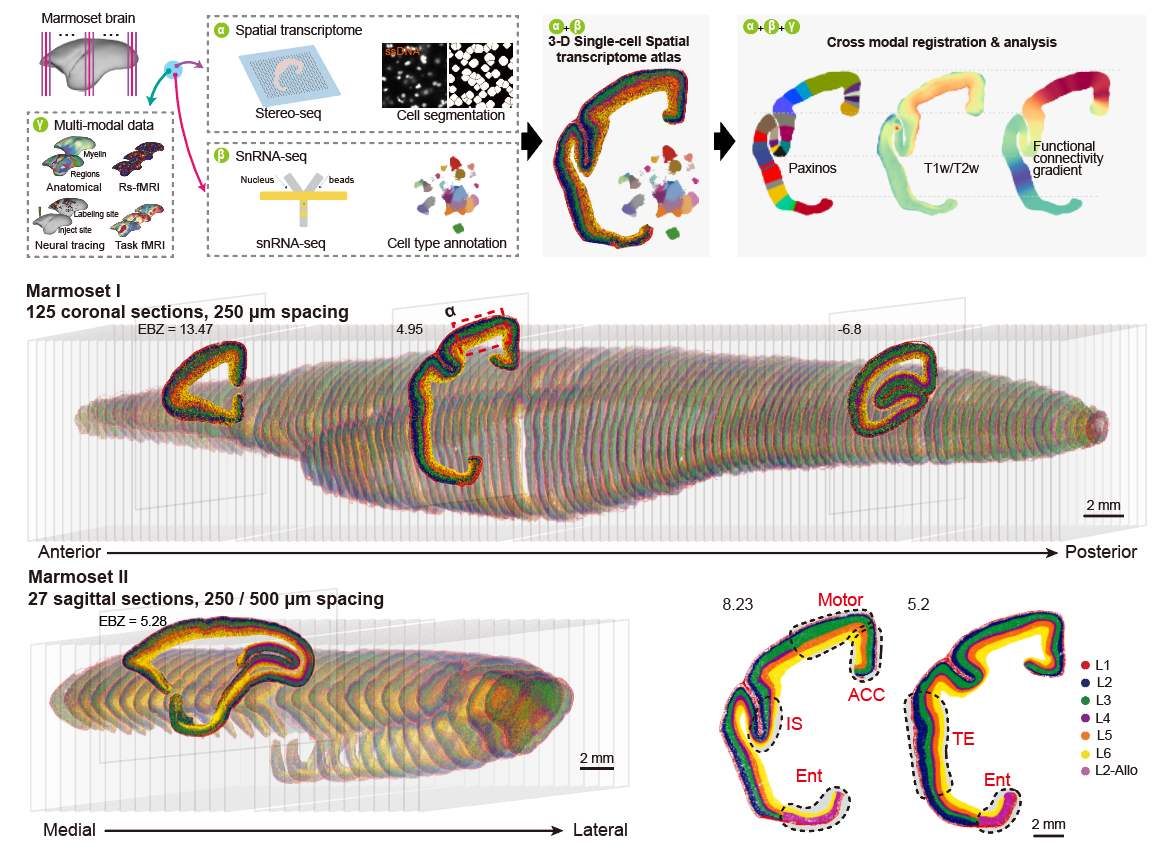 Single-cell spatial transcriptomic atlas of the marmoset cortex.
Single-cell spatial transcriptomic atlas of the marmoset cortex.
Analysis revealed the key finding: two opposing molecular gradients that define the dominant axis of cortical organization. One radiates from allocortical regions, the evolutionarily ancient areas involved in smell and deep memory, while the other originates from the primary sensory cortices that handle the raw signals of seeing, hearing, and touch. The association cortex, which integrates this information to support reading, decision-making, and abstract thought, sits at their intersection.
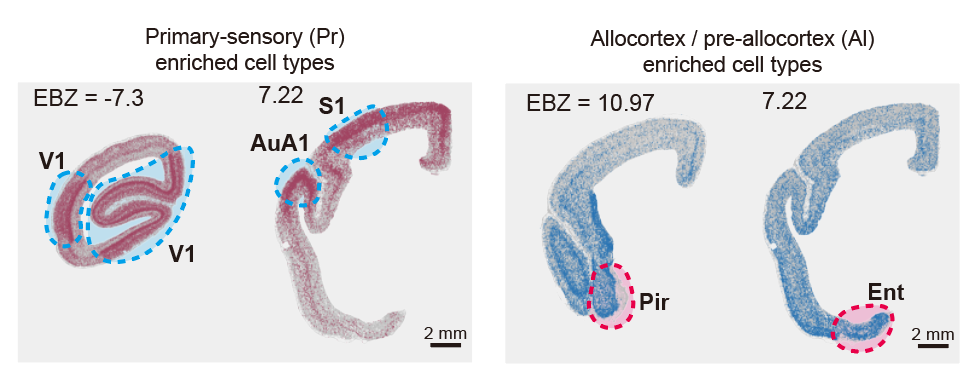
Spatial maps of cell types enriched in primary-sensory (Pr) and allocortical/per-iallocortical (Al) regions.
The team quantified this architecture using a Pr-Al index that captured roughly 40 percent of total variance in cortical cell composition, and the same pattern was conserved across humans, macaques, marmosets, and mice. Rather than one hypothesis being correct and the other wrong, both allocortical regions and primary sensory areas serve as genuine organizational anchors at opposite ends of a single continuous axis. These gradients are present at birth but undergo substantial postnatal refinement, with gradient segregation increasing nearly sixfold from newborns to adults, consistent with ongoing maturation shaped by sensory experience.
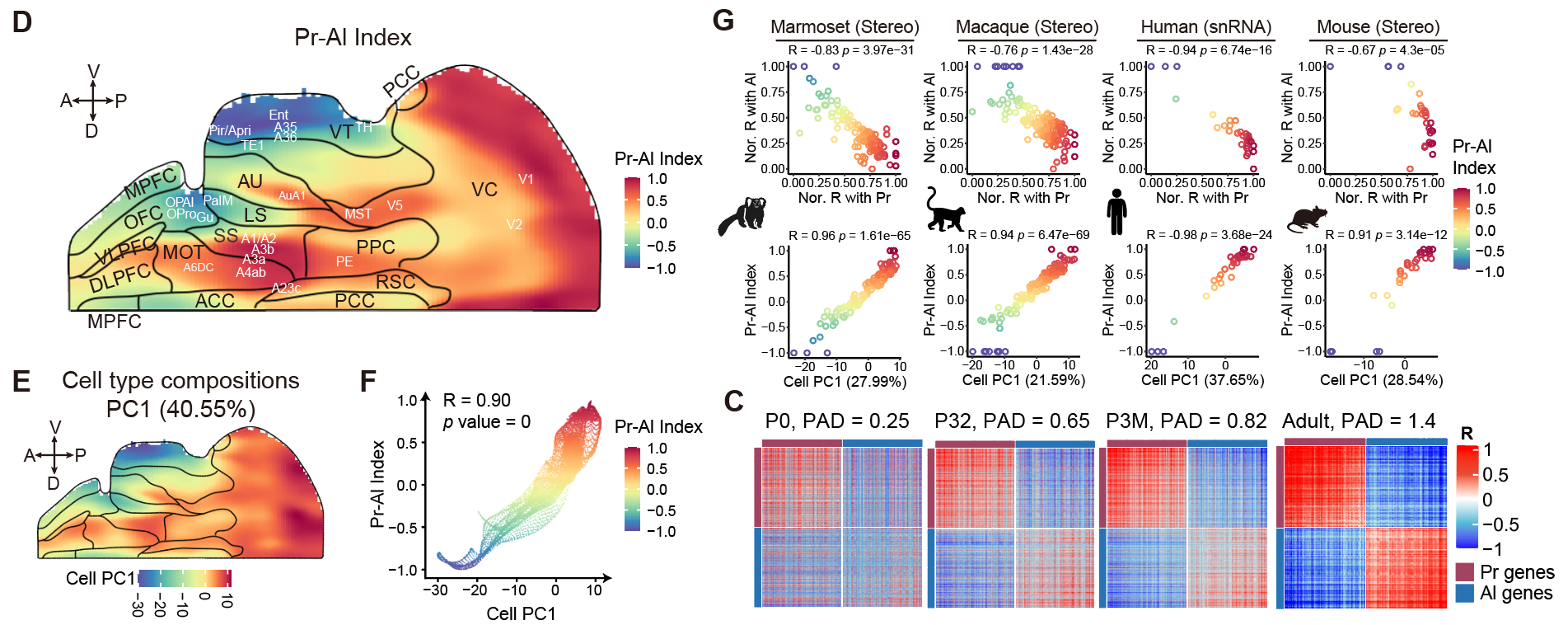
Spatial transcriptomic mapping across species and developmental stages reveals two opposing cortical gradients that sharpen with age, reconciling long-standing theories of cortical organization.
The gradients extend beyond the cortex. Cortical gradient-associated gene sets showed strongly anticorrelated patterns in the thalamus, aligning with thalamocortical connectivity confirmed by retrograde tracing,indicating that the two did not evolve independently, but are regulated by a shared conserved molecular program and form the core organizational framework of the thalamocortical system. At the zone where the two gradients converge, the putative default mode network and frontal pole shared similar molecular signatures despite weak functional connectivity, suggesting that a shared molecular identity may have evolved before the strong network integration seen in humans.
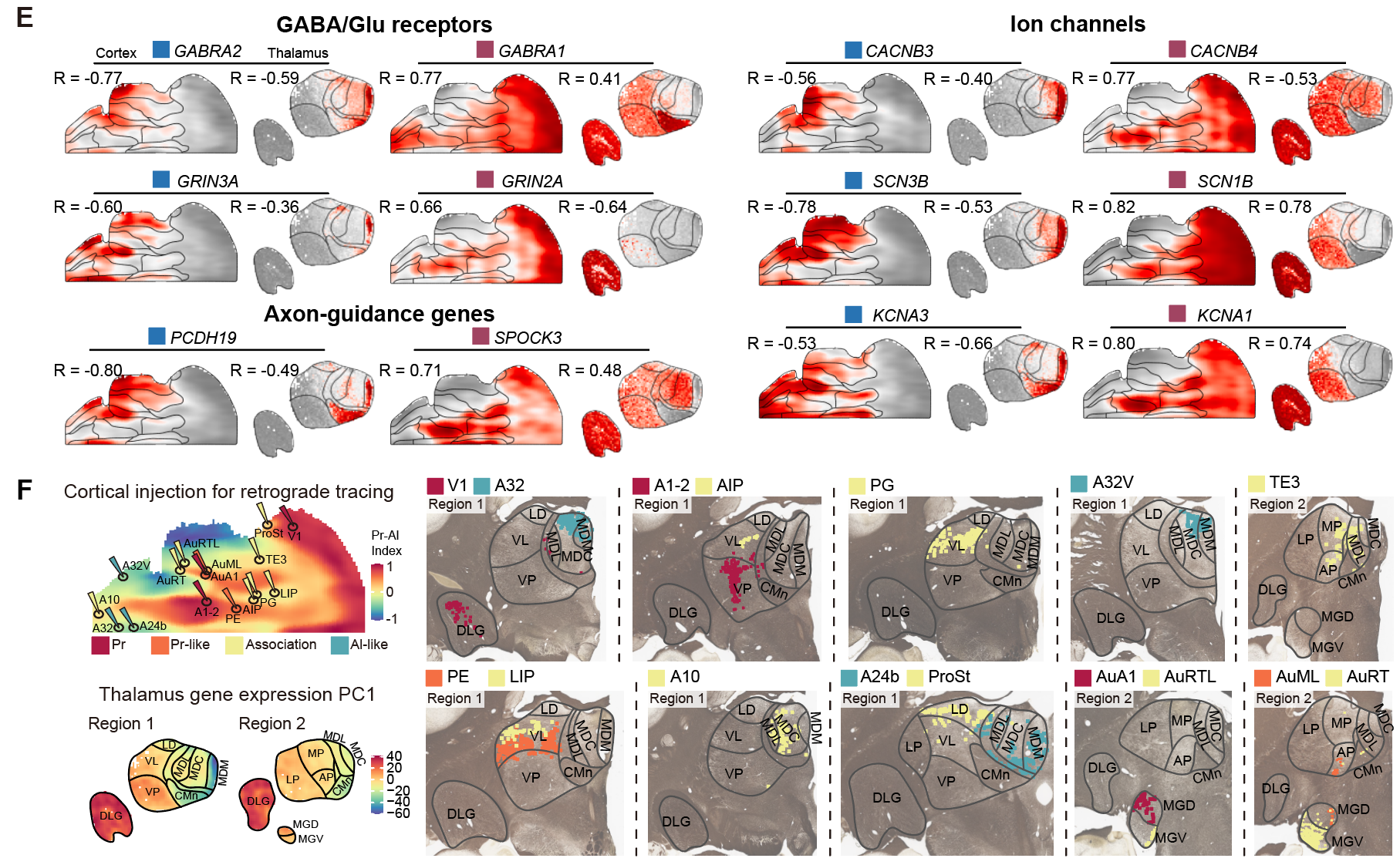
Cortical gradients are mirrored in thalamic gene expression and connectivity patterns.
Perhaps most surprisingly, although macaques are phylogenetically closer to humans, the marmoset auditory cortex shared significantly more molecular features with the human auditory cortex. Integrative analysis identified 984 differentially expressed genes shared between humans and marmosets but not macaques, far exceeding the 269 shared exclusively between humans and macaques. Because both humans and marmosets exhibit sophisticated vocal behaviors including turn-taking and vocal learning, this convergence may reflect shared selective pressures for complex auditory processing, positioning the marmoset as a potentially valuable model for studying the molecular basis of vocal communication.
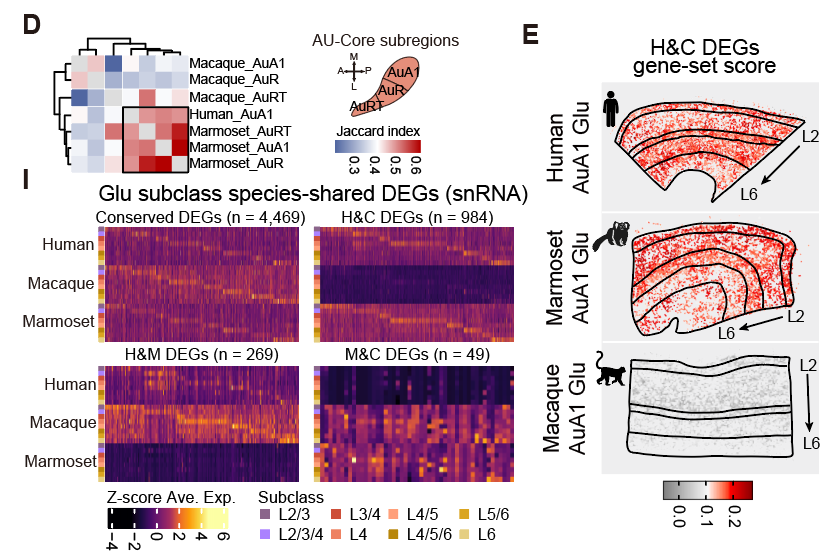
Cross-species comparison of auditory cortex reveals greater molecular similarity between humans and marmosets than macaques, positioning the marmoset as a key model for studying language evolution.
All animal studies and procedures were approved by the Animal Care and Use Committee of CEBSIT, Chinese Academy of Sciences. All data have been deposited in MCCSTA databases at CNGBdb, and analysis code is publicly available.
This research can be accessed at https://doi.org/10.1126/science.aea2673.



